Using a new non-toxic imaging method, researchers watched living human embryos divide in real time and discovered that chromosome separation errors linked to mosaicism can arise even at the blastocyst stage, showing that mosaicism is a dynamic process rather than one confined to early development.
Aneuploidy, an abnormal number of chromosomes, is a leading cause of implantation failure, miscarriage, and abnormal embryo development.
If the error happens during the formation of the egg or sperm (meiosis), every cell in the embryo is affected. But if it happens after fertilization (mitosis), only some cells are affected, creating a mix of normal and abnormal cells known as mosaicism.
You can see the difference below, with two examples of a mitotic error that occur during the two-cell or four-cell stage.
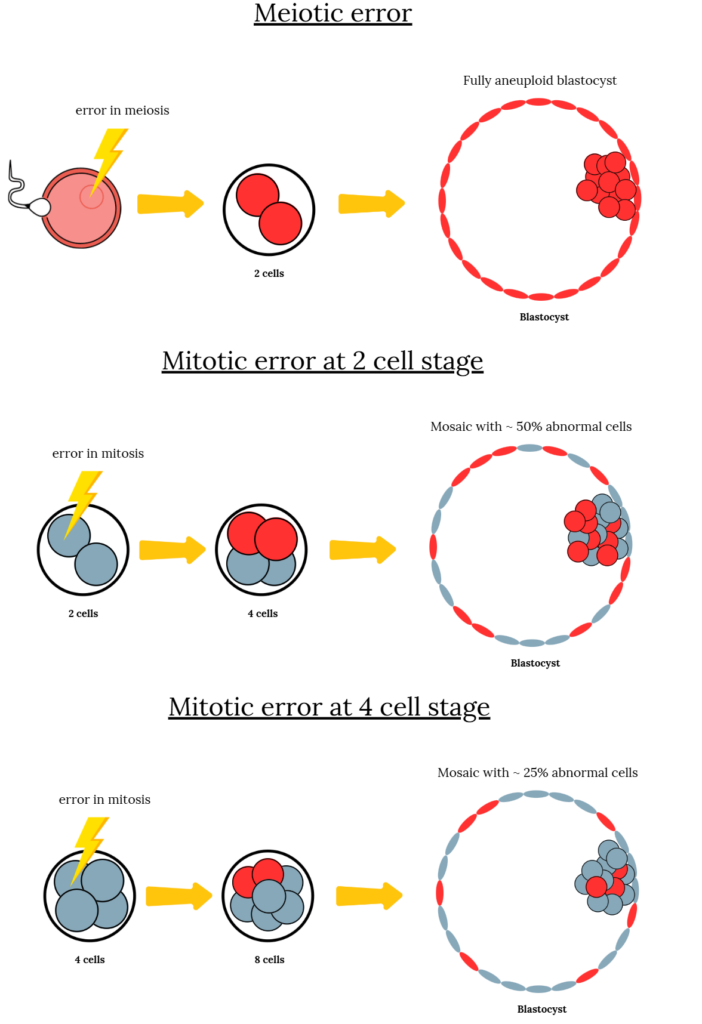
While studies have shown that mosaicism can occur during the cleavage stage (day 2-3), it’s less clear whether new errors continue to appear during the blastocyst stage (day 5-7). In a new study by Abdelbaki et al. (2025), researchers developed a gentle, non-toxic imaging method that allowed them to label DNA and watch living human blastocysts divide in real time, revealing how new chromosome errors can still form just before implantation.
🔗 Original studies are referenced in this post or within the linked Remembryo posts.
💡 Reminder: Terms underlined with a dotted black line are linked to glossary entries. Clicking these does not count toward your paywall limit.
How do chromosomal errors occur?
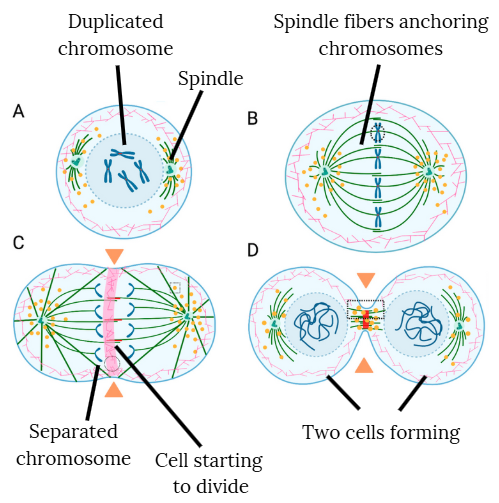
- When cells divide in an embryo, each cell must first copy its DNA so that both new cells receive the full set of 46 chromosomes. To do this, the chromosomes are duplicated, forming an X shape made of two identical copies joined together (panel A).
- These duplicated chromosomes line up in the middle of the cell (panel B), where a structure called the spindle uses fibers to attach to each duplicated chromosome, in preparation for pulling them apart into single chromosomes.
- In panel C, you can see the duplicate chromosomes being separated into single chromosomes.
- As the cell divides into two (panel D), each new cell receives one complete set of chromosomes.
- If the spindle doesn’t anchor or pull correctly, or if the chromosomes are misaligned or lagging behind as the spindle pulls, this can lead to chromosome segregation (separation) errors. This can lead to some cells ending up with too many or too few chromosomes, also known as aneuploidy.
Want more background or details? Check out my post Chromosomal errors in IVF: What is aneuploidy and what causes it?
Chromosomal separation errors seen as blastocysts’ cells divide
By using 13 human blastocysts and 17 mouse blastocysts, the researchers visualized nearly 500 cell divisions to see if any chromosome separation errors occurred. Roughly 8% of human divisions showed misalignment or lagging chromosomes (compared with 4% in mice). Rare events included multipolar divisions (1% of divisions) and mitotic slippage, where a cell exits division without properly separating, potentially creating tetraploid cells with twice the normal amount of DNA.
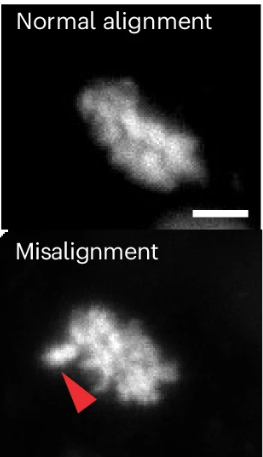
They also saw micronuclei, tiny structures that contain DNA that form when chromosomes don’t separate correctly. Most of these stayed outside the main nucleus and were passed to one daughter cell. The affected cells kept dividing, suggesting that embryos can tolerate these errors rather than removing them.
The researchers also found that most cells with chromosome separation errors stayed in the trophectoderm, with only rare movement into the ICM, suggesting that most errors occur in placental rather than embryonic cells.
Conclusions
This study highlights the work of a new imaging method that lets scientists watch chromosomes and cells in living human embryos. This technique could help future studies explore how embryos develop and how chromosomal errors arise.
Using this technique, the researchers showed that chromosomal separation errors occur during the blastocyst stage, including misaligned or lagging chromosomes, multipolar divisions, and micronuclei. While they didn’t show that aneuploidy actually occurred by PGT-A or another genetic test, these types of chromosome separation errors are what directly cause aneuploidy.
They also found that cells with these errors persisted in the trophectoderm and continued dividing instead of dying, as would typically happen in other cell types, suggesting that trophectoderm cells may have mechanisms that allow them to tolerate these errors.
These results show that the developing blastocyst is dynamic and can continue to acquire abnormalities as its cells divide. Not all mosaic abnormalities detected by PGT-A therefore occur during the cleavage stage, and some may develop later as the embryo continues to develop. This isn’t really surprising, but to my knowledge, it’s the first study to directly visualize it.
This means that any PGT-A result — whether euploid, mosaic, or aneuploid — represents only a snapshot of the embryo’s genetic state at the moment of biopsy, and that new mosaic changes can continue to develop as the embryo grows.
But what does this mean clinically?
- The study didn’t confirm aneuploidy through PGT-A. Instead, it detected markers of chromosomal instability. These are events that could lead to aneuploidy but don’t always result in it.
- These late-stage errors may represent the earliest steps toward mosaicism. For an error to be detected as mosaic by conventional PGT-A, the affected cell must divide many times. Errors that occur early, during the cleavage stage, have more time to multiply and can form high-level mosaics that more strongly reduce success rates. Errors that appear later, at the blastocyst stage, likely don’t have enough time to divide before transfer to influence outcomes (studies have shown that low level mosaics can have comparable outcomes to euploids).
- The fact that conventional PGT-A would miss these late errors probably isn’t clinically significant. High-level mosaics, which are more relevant in reducing success rates, likely form earlier in development and would already be detectable by PGT-A at the blastocyst stage.
- This study doesn’t change that PGT-A is still a valuable tool for selecting embryos, since PGT-A results are linked to different success rates. The issues with PGT-A mainly stem from potentially discarding embryos that are viable. This study did find that errors were confined to the trophectoderm, but they didn’t really assess the ICM, which develops into the fetus. Future studies will hopefully look at this using this new technology.
Still, these findings fit with what other research has shown. Other studies that use more sensitive PGT-A methods, listed below, have shown that mosaicism is relatively common in blastocysts, sometimes occurring at levels too low to be detected by standard PGT-A methods. This new study supports those findings and may explain how these very low levels of mosaicism develop.
Want to read more about mosaic embryos and PGT-A?
Aneuploid embryos, with an abnormal number of chromosomes, are a major cause of miscarriages, often linked to maternal age and errors in meiosis. In this post, we'll explore what chromosomes and aneuploidy are, how meiosis errors cause aneuploidy, and the factors contributing to these errors, including age, genetics and mitochondrial dysfunction. Read more.
This post covers mosaic embryos, including their definition, frequency after PGT-A, mosaic embryo levels (low level, high level), mosaic embryo types (whole chromosome, segmental), current guidelines for transfer, potential for self-correction, and success rates. Read more.
Researchers in a 2021 study investigated "self-correction" in a stem cell-based model for mosaic embryos and found that aneuploid cells in a mosaic are more likely to change into trophectoderm cells or die, while euploid cells accumulate in the ICM. Read more.
A 2025 study re-analyzed embryos that tested as mosaic by NGS-based PGT-A using SNP karyotyping and found that one-third of those "mosaics" were actually uniformly aneuploid. Read more.
A 2024 study performed a more sensitive type of PGT-A by analyzing single cells, finding that nearly all blastocysts and fetal tissue contain some level of mosaicism. Read more.
Reference
If you liked this post and want to support what I do, please consider a paid subscription, Patreon or donate through PayPal!

About Embryoman
Embryoman (Sean Lauber) is a former embryologist and the founder of Remembryo, an IVF research and fertility education website. After working in an IVF lab in the US, he returned to Canada and now focuses on making fertility research more accessible. He holds a Master’s in Immunology and launched Remembryo in 2018 to help patients and professionals make sense of IVF research. Sean shares weekly study updates on Facebook, Instagram, and Reddit regularly. He also answers questions on Reddit or in his private Facebook group.














